Fragment Screening & Fragment-Based Lead Discovery
In the traditional high-throughput screening method (HTS), a library of drug-like compounds, which may consist of hundreds of thousands or even millions of compounds, are screened against the protein target. Such screening may identify several active compounds that may bind to the drug target and inhibit its activity. However, a more efficient method that requires a much smaller pool of compounds is fragment-based drug discovery (FBDD). In this approach, screening is run using a library of small molecule fragments (molecular weight not larger than 300 Da), usually called a "fragment library." Since the molecules used in this case are smaller and less complex than drug-like molecules, the probability of finding a molecule that will fit into the protein binding site is higher. Using fragments, it is also possible to better sample the chemical space. For choosing compounds for the library, the Rule of Three has been suggested instead of Lipinski Rule of 5:
Fragment-based drug design allows the size of the compound library to be kept much smaller (it could be just a few hundred rather than hundreds of thousands), which is much easier to handle practically. Small fragments may easily bind to various active site pockets, which are not accessible for larger, drug-like molecules. The aim is to find weak but efficient binders (high binding energy for the atoms involved in the interactions). Bound fragments may subsequently be grown into more potent compounds with drug-like properties in a process called hit expansion. This is shown schematically in the image below:
- MW 100-300
- LogP ≤ 3.0
- H-Bond Acceptors ≤ 3
- H-Bond Donors ≤ 3
- Rotatable bonds ≤ 3
- Polar Surface Area ≤ 60 Å2
Fragment-based drug design allows the size of the compound library to be kept much smaller (it could be just a few hundred rather than hundreds of thousands), which is much easier to handle practically. Small fragments may easily bind to various active site pockets, which are not accessible for larger, drug-like molecules. The aim is to find weak but efficient binders (high binding energy for the atoms involved in the interactions). Bound fragments may subsequently be grown into more potent compounds with drug-like properties in a process called hit expansion. This is shown schematically in the image below:
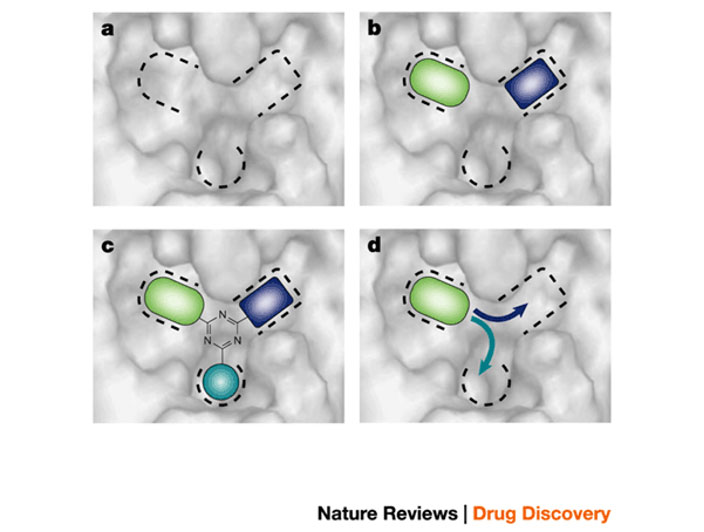
Schematic image showing how fragment-based lead discovery is applied to a case of a binding site comprising three potential binding pockets
a ) Screening locates molecular fragments that bind into one (d), two (b), or all three pockets (c).
b) A lead compound is designed by organizing three fragments around a core template, as shown in (c)
c ) It could be grown out from a single fragment, as shown in (d)
Blundell et al. (2002), Nature Rev Drug Discov. 1, 45-54.
b) A lead compound is designed by organizing three fragments around a core template, as shown in (c)
c ) It could be grown out from a single fragment, as shown in (d)
Blundell et al. (2002), Nature Rev Drug Discov. 1, 45-54.
The advantage of the fragment-based approach is the ability to systematically grow the bound fragment during a hit expansion process. This gives better control of the parameters of the molecules (such as solubility, toxicity, etc.). Currently, many examples, starting from identifying the first fragment hits, followed by hit expansion, lead generation, and optimization, made it all the way to drug candidates and clinical trials. The combination of fragment-based drug discovery and structure-based drug design is superior to old drug discovery methods (P.J. Hajduk & J. Greer, 2007).