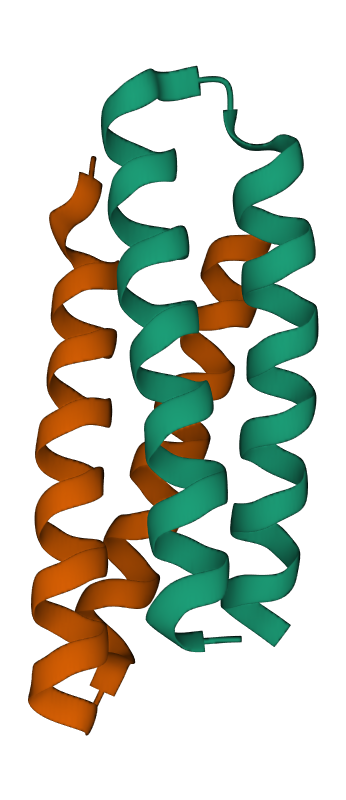Protein Domains, Folds, and Motifs
In protein structures, helices and strands may be connected and combined in many ways to form a domain. However, from known protein three-dimensional structures, we have learned that there is only a limited number of ways by which secondary structure elements are combined in proteins. These different ways are called folds (or Topology according to CATH database classification). Therefore, we will examine typical examples of connectivity and packing secondary structure elements within 3D structures. This type of analysis simplifies the comparison of protein structures and reveals similarities and differences, opening the way for structure classification.
Defining a domain
A domain may be characterized by the following:
1- Spatially separated unit of the protein structure
2- Often has sequence and/or structural resemblance to other protein structures or domains.
3- Often has a specific function associated with it.
Some proteins (examples include triose phosphate isomerase (TIM), plastocyanin, and hemoglobin) contain one single domain. However, a large number of proteins are multidomain proteins that may contain two or more domains. Here, we start by analyzing single domains, normally found within multidomain proteins.
The helix bundle domain
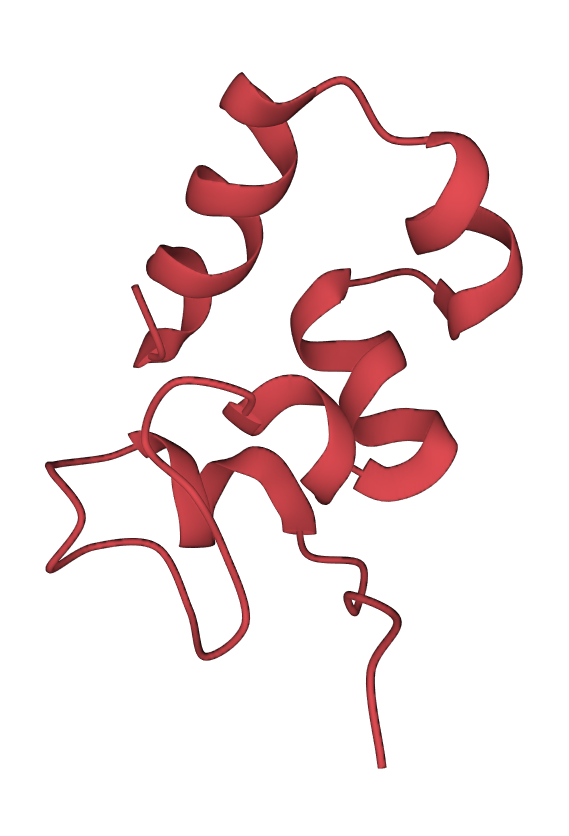
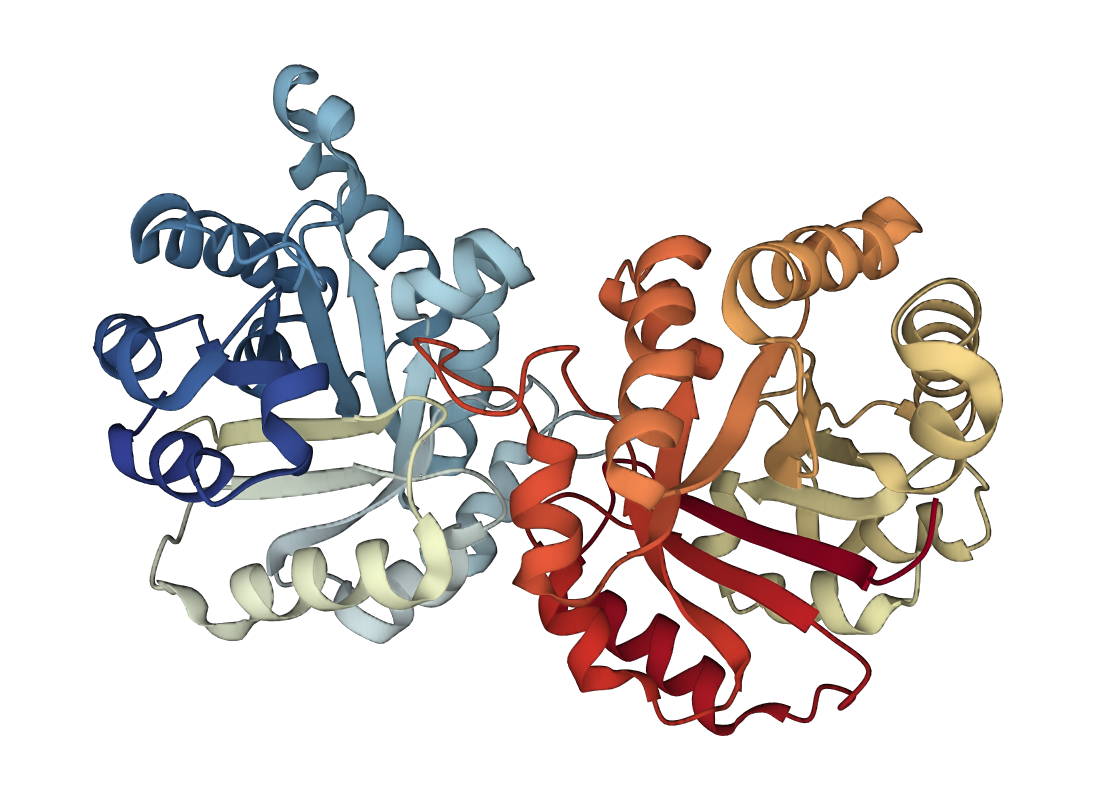
One of the common domains is the helix bundle domain (the images below show two different bundles). Helix bundles are often found as separate domains within larger, multidomain proteins. On the left image is the crystal structure of a De Novo designed protein (PDB 1MFT). In this case, the helices within the bundle are connected by short loops and packed to form a hydrophobic core in the middle of the bundle. On the right is a solution structure of the headpiece domain of chicken villin (PDB 1QQV). Here the arrangement of the helices is very different, and the loops are much longer. Clicking the images will take you to the RCSB PDB 3D-viewer, where you can rotate the structures to get a better impression of how they are built:
Defining a domain
A domain may be characterized by the following:
1- Spatially separated unit of the protein structure
2- Often has sequence and/or structural resemblance to other protein structures or domains.
3- Often has a specific function associated with it.
Some proteins (examples include triose phosphate isomerase (TIM), plastocyanin, and hemoglobin) contain one single domain. However, a large number of proteins are multidomain proteins that may contain two or more domains. Here, we start by analyzing single domains, normally found within multidomain proteins.
The helix bundle domain
One of the common domains is the helix bundle domain (the images below show two different bundles). Helix bundles are often found as separate domains within larger, multidomain proteins. On the left image is the crystal structure of a De Novo designed protein (PDB 1MFT). In this case, the helices within the bundle are connected by short loops and packed to form a hydrophobic core in the middle of the bundle. On the right is a solution structure of the headpiece domain of chicken villin (PDB 1QQV). Here the arrangement of the helices is very different, and the loops are much longer. Clicking the images will take you to the RCSB PDB 3D-viewer, where you can rotate the structures to get a better impression of how they are built:
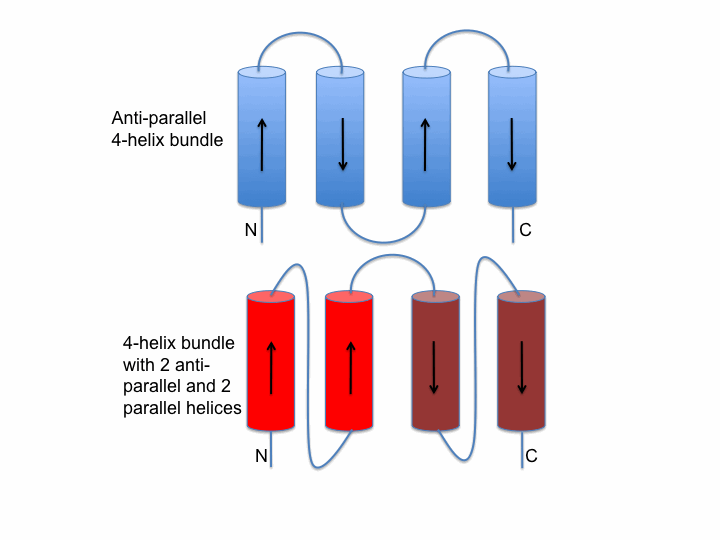
Helix bundle packing
The image on the left shows two schematic examples of connectivity between helices in a bundle. The upper packing (blue) shows an antiparallel bundle. In contrast, the lower bundle (red) shows an example where two helices are parallel to each other and antiparallel to the second pair.
The image on the left shows two schematic examples of connectivity between helices in a bundle. The upper packing (blue) shows an antiparallel bundle. In contrast, the lower bundle (red) shows an example where two helices are parallel to each other and antiparallel to the second pair.
Connecting strands and helices: The Rossmann fold - the coenzyme binding domain
The strands within parallel and antiparallel β-sheets may be connected by structural elements like loops, helices, or coil regions (regions without a defined secondary structure). As described earlier, loops in the hairpin motif make the simplest and most common connectivity between strands.
An example of helix-strand connectivity is provided by the Rossmann fold domain, named after Michael G. Rossmann, a protein crystallographer who solved the structure of lactate dehydrogenase (LDH). The Rossmann fold is the only protein fold named after the person who discovered it. This domain type is widespread and can be found in many multidomain proteins involved in binding nucleotide cofactors like NADH, FAD, FMN, as well as ATP and GTP. An excellent discussion of details and the history of the Rossmann fold can be found on Proteopedia.
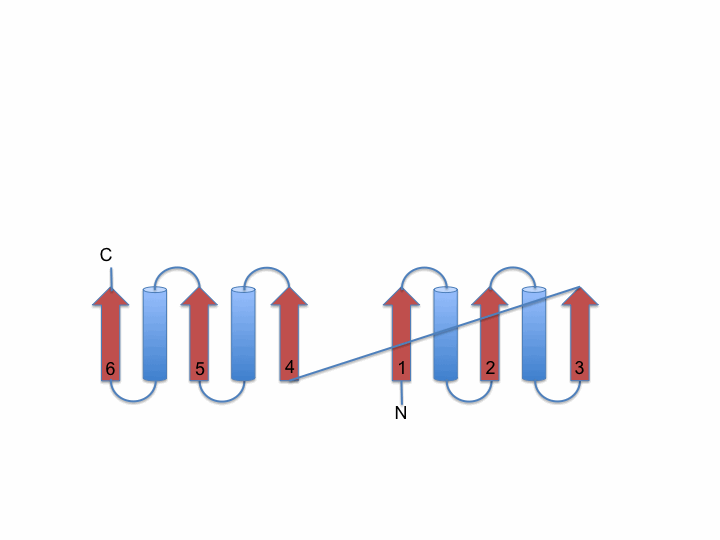
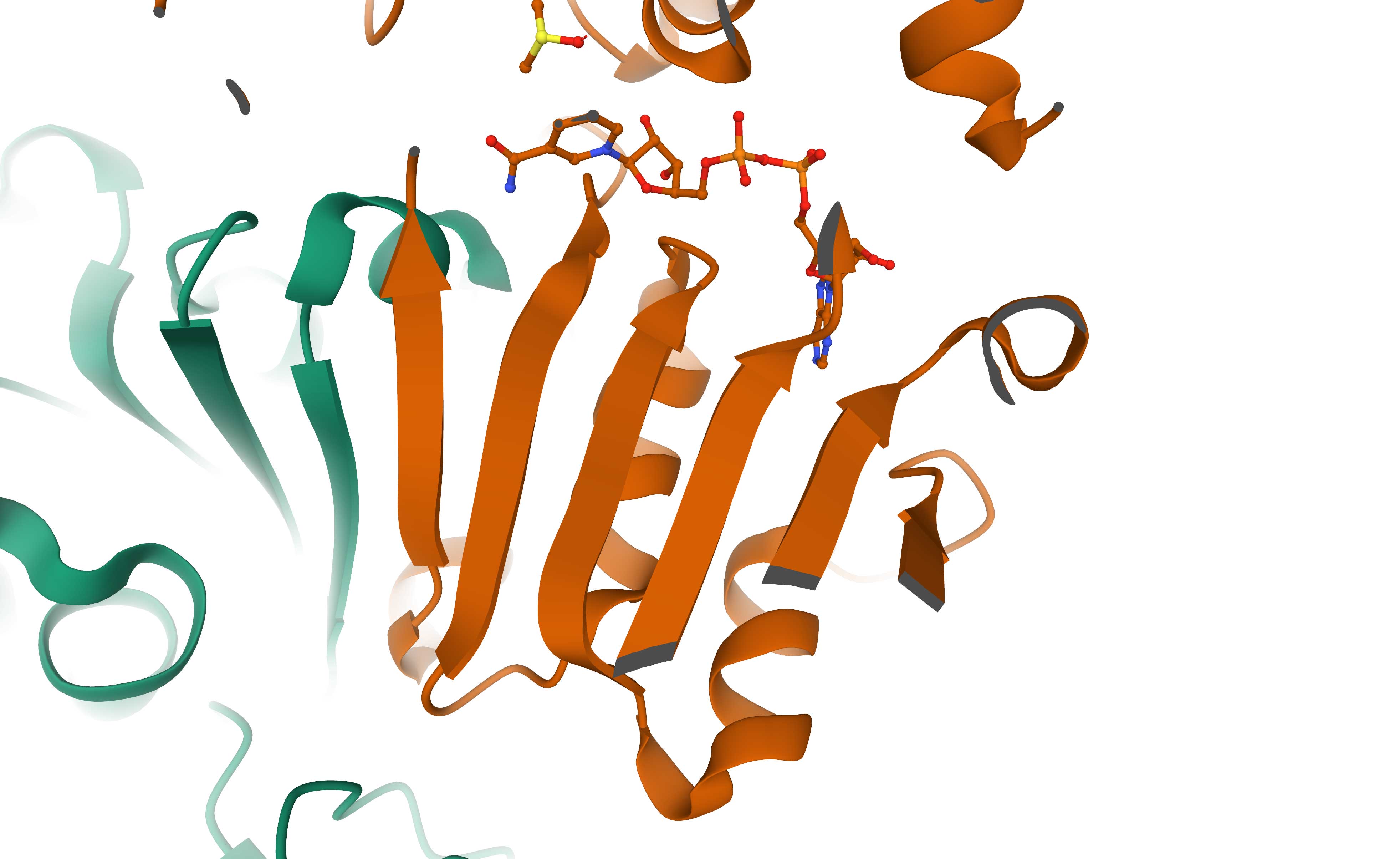
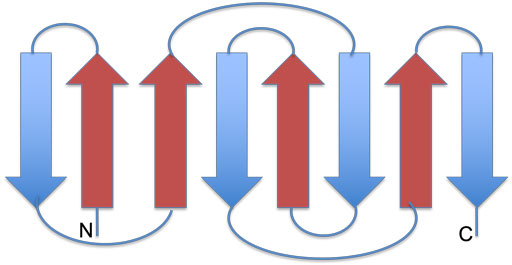
The left image below shows a schematic representation of the Rossmann fold. It consists of a parallel 6-stranded β-sheet flanked by α-helices. The right image shows the 6-stranded parallel Rossmann fold β-sheet of the enzyme liver alcohol dehydrogenase (LDH). The NADH molecule bound at the top of the β-sheet is shown as a stick model (PDB 2OHX). The parallel β-sheet (in yellow) is flanked by α-helices on both sides of its plane. The helices which should be on top of the sheet are removed here for clarity.
The strands within parallel and antiparallel β-sheets may be connected by structural elements like loops, helices, or coil regions (regions without a defined secondary structure). As described earlier, loops in the hairpin motif make the simplest and most common connectivity between strands.
An example of helix-strand connectivity is provided by the Rossmann fold domain, named after Michael G. Rossmann, a protein crystallographer who solved the structure of lactate dehydrogenase (LDH). The Rossmann fold is the only protein fold named after the person who discovered it. This domain type is widespread and can be found in many multidomain proteins involved in binding nucleotide cofactors like NADH, FAD, FMN, as well as ATP and GTP. An excellent discussion of details and the history of the Rossmann fold can be found on Proteopedia.
The left image below shows a schematic representation of the Rossmann fold. It consists of a parallel 6-stranded β-sheet flanked by α-helices. The right image shows the 6-stranded parallel Rossmann fold β-sheet of the enzyme liver alcohol dehydrogenase (LDH). The NADH molecule bound at the top of the β-sheet is shown as a stick model (PDB 2OHX). The parallel β-sheet (in yellow) is flanked by α-helices on both sides of its plane. The helices which should be on top of the sheet are removed here for clarity.
The TIM barrel domain
Triose phosphate IsoMerase (TIM) is a single-domain protein that is also the first representative of the TIM barrel fold. This fold is also widespread and found in many proteins. Here the strands of the β-sheet are parallel and connected by loops and helices. Details of the mechanism and function of this protein can be found on Proteopedia. On the left image is a schematic presentation of the fold, while on the right is a ribbon presentation of the three-dimensional structure of triose phosphate isomerase dimer. Again, clicking on the image will open a graphics window where you can rotate the structure to get a better impression of the fold.
More on strand connectivity
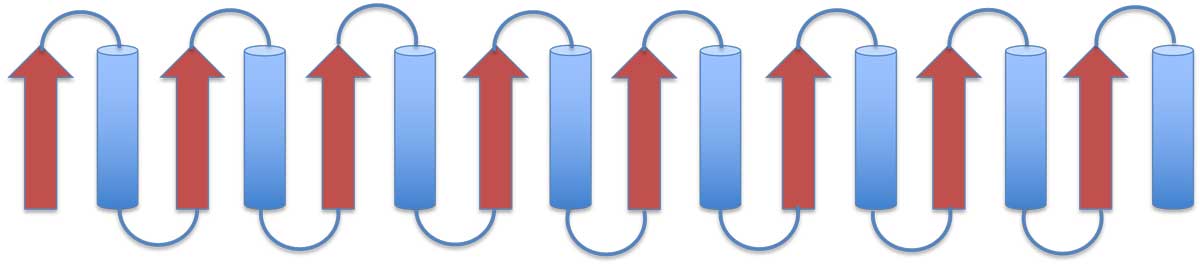
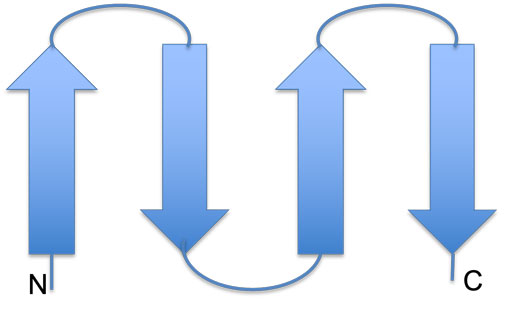
Examples of connectivity in antiparallel sheets are shown below. On the left, two hairpins are connected into an antiparallel β-sheet, and on the right is the Greek-key motif:
Examples of connectivity in antiparallel sheets are shown below. On the left, two hairpins are connected into an antiparallel β-sheet, and on the right is the Greek-key motif:

The following image shows the protein plastocyanin (PDB 1bxu), which predominantly contains β–strands and coiled regions. A couple of short helical turns are also shown in red.
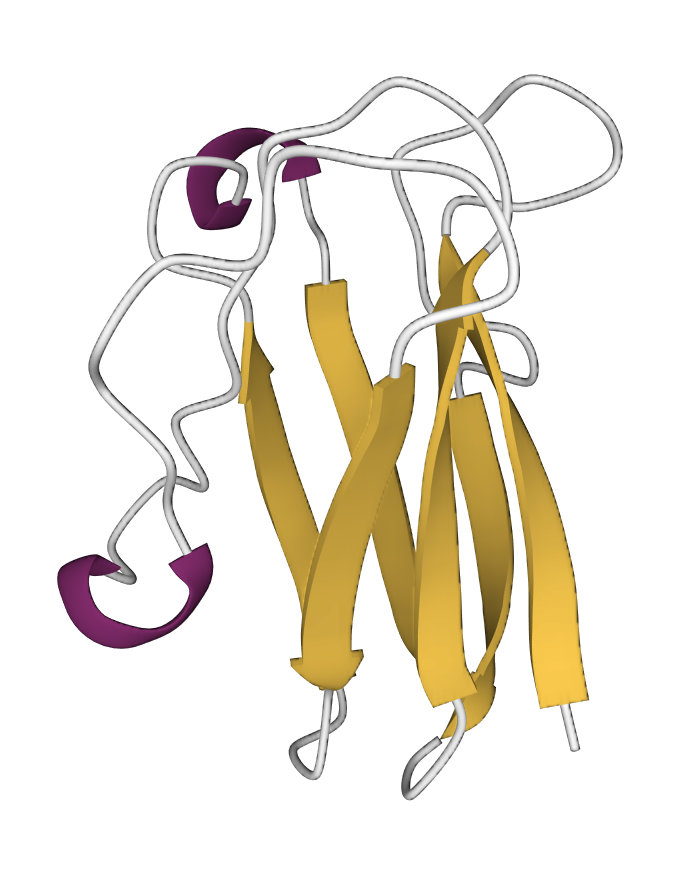
These are just some examples of protein domains and single-domain proteins with different types of folds and structural motifs demonstrating various types of connectivity between secondary structure elements. There is, of course, a much larger number of other folds (around 1300). In the following section, we will examine the classification of domains and folds, a way to bring order into the system.